diff --git a/Plugins/org.mitk.gui.qt.diffusionimaging.fiberfox/src/internal/QmitkFiberGenerationViewControls.ui b/Plugins/org.mitk.gui.qt.diffusionimaging.fiberfox/src/internal/QmitkFiberGenerationViewControls.ui
index 29f96f5..3536f99 100644
--- a/Plugins/org.mitk.gui.qt.diffusionimaging.fiberfox/src/internal/QmitkFiberGenerationViewControls.ui
+++ b/Plugins/org.mitk.gui.qt.diffusionimaging.fiberfox/src/internal/QmitkFiberGenerationViewControls.ui
@@ -1,2012 +1,2012 @@
QmitkFiberGenerationViewControls
0
0
413
1461
Form
-
Save Parameters
:/QmitkDiffusionImaging/general_icons/download.ico:/QmitkDiffusionImaging/general_icons/download.ico
-
Load Parameters
:/QmitkDiffusionImaging/general_icons/upload.ico:/QmitkDiffusionImaging/general_icons/upload.ico
-
Qt::Vertical
20
40
-
- 0
+ 1
0
0
381
731
Manual Fiber Design
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
false
30
30
Draw elliptical fiducial.
:/QmitkFiberfox/circle.png:/QmitkFiberfox/circle.png
32
32
false
true
-
false
30
30
Flip fiber waypoints of selcted fiducial around one axis.
:/QmitkFiberfox/refresh.xpm:/QmitkFiberfox/refresh.xpm
32
32
false
true
-
Qt::Horizontal
40
20
-
color: rgb(255, 0, 0);
Please select an image or an existing fiber bundle to draw the fiber fiducials. If you can't provide a suitable image, generate one using the Fiberfox View.
Qt::AutoText
Qt::AlignJustify|Qt::AlignVCenter
true
-
QGroupBox {
background-color: transparent;
}
Fiber Options
6
6
6
6
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
Tension:
false
-
Fiber Sampling:
false
-
3
-1.000000000000000
1.000000000000000
0.100000000000000
0.000000000000000
-
3
-1.000000000000000
1.000000000000000
0.100000000000000
0.000000000000000
-
Bias:
false
-
Continuity:
false
-
3
-1.000000000000000
1.000000000000000
0.100000000000000
0.000000000000000
-
Distance of fiber sampling points (in mm)
1
0.100000000000000
0.100000000000000
1.000000000000000
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
6
-
#Fibers:
false
-
Specify number of fibers to generate for the selected bundle.
1
1000000
100
100
-
false
QCommandLinkButton:disabled {
border: none;
}
Generate Fibers
:/QmitkDiffusionImaging/general_icons/right.ico:/QmitkDiffusionImaging/general_icons/right.ico
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
Select fiber distribution inside of the fiducials.
-
Uniform
-
Gaussian
-
Fiber Distribution:
false
-
Variance of the gaussian
3
0.001000000000000
10.000000000000000
0.010000000000000
0.100000000000000
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
Disable to only generate fibers if "Generate Fibers" button is pressed.
Real Time Fibers
true
-
Disable to only generate fibers if "Generate Fibers" button is pressed.
Advanced Options
false
-
QGroupBox {
background-color: transparent;
}
Fiducial Options
6
6
6
6
-
Fiducial Attributes
6
6
6
6
-
Length of fiducial axis 2
2
0.000000000000000
9999.000000000000000
0.100000000000000
-
Axis 1:
false
-
Position X:
false
-
World Position in mm
2
-99999.000000000000000
99999.000000000000000
0.100000000000000
-
World Position in mm
2
-360.000000000000000
360.000000000000000
0.100000000000000
-
World Position in mm
2
-360.000000000000000
360.000000000000000
0.100000000000000
-
Length of fiducial axis 1
2
0.000000000000000
9999.000000000000000
0.100000000000000
-
Position Z:
false
-
Position Y:
false
-
Axis 2:
false
-
Twist:
false
-
Twist in degree
2
-180.000000000000000
180.000000000000000
0.100000000000000
-
All fiducials are treated as circles with the same radius as the first fiducial.
Use Constant Fiducial Radius
false
-
false
Align selected fiducials with voxel grid. Shifts selected fiducials to nearest voxel center.
QCommandLinkButton:disabled {
border: none;
}
Align With Grid
:/QmitkDiffusionImaging/general_icons/right.ico:/QmitkDiffusionImaging/general_icons/right.ico
0
0
395
277
Random Phantom
-
Volume Size:
false
-
Curvyness:
false
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
- Specify number of fibers to generate for the selected bundle.
+ Max. fiber twist per step (in degree)
0
90
10
30
-
- Specify number of fibers to generate for the selected bundle.
+ Min. fiber twist per step (in degree)
0
90
10
15
-
Step Size:
false
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
- Specify number of fibers to generate for the selected bundle.
+ Min. distance between fiducials (in mm)
1
100
1
15
-
- Specify number of fibers to generate for the selected bundle.
+ Max. distance between fiducials (in mm)
1
100
1
30
-
Streamline Density:
false
-
- Specify number of fibers to generate for the selected bundle.
+ Number of generated bundles
1
1000
1
50
-
Start Radii:
false
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
- Specify number of fibers to generate for the selected bundle.
+ Min. rotation between steps (in degree)
1
90
1
5
-
- Specify number of fibers to generate for the selected bundle.
+ Max. rotation between steps (in degree)
1
90
1
45
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
Specify number of fibers to generate for the selected bundle.
1
1000
1
250
-
Specify number of fibers to generate for the selected bundle.
1
1000
1
250
-
Specify number of fibers to generate for the selected bundle.
1
1000
1
250
-
Twist:
false
-
#Bundles:
false
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
- Specify number of fibers to generate for the selected bundle.
+ Bundle radius
1
300
1
5
-
- Specify number of fibers to generate for the selected bundle.
+ Bundle radius
1
300
1
25
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
- Specify number of fibers to generate for the selected bundle.
+ <html><head/><body><p>Max. fibers per cm<span style=" vertical-align:super;">2</span></p></body></html>
1
10000
10
200
-
- Specify number of fibers to generate for the selected bundle.
+ <html><head/><body><p>Min. fibers per cm<span style=" vertical-align:super;">2</span></p></body></html>
1
10000
10
50
-
true
QCommandLinkButton:disabled {
border: none;
}
Generate
:/QmitkDiffusionImaging/general_icons/right.ico:/QmitkDiffusionImaging/general_icons/right.ico
0
0
395
283
Operations
-
QFrame::NoFrame
QFrame::Raised
0
0
0
0
-
Y
false
-
Rotation angle (in degree) around x-axis.
2
-360.000000000000000
360.000000000000000
0.100000000000000
-
Axis:
false
-
Rotation angle (in degree) around y-axis.
2
-360.000000000000000
360.000000000000000
0.100000000000000
-
Translation:
false
-
Translation (in mm) in direction of the z-axis.
2
-1000.000000000000000
1000.000000000000000
0.100000000000000
-
Translation (in mm) in direction of the y-axis.
2
-1000.000000000000000
1000.000000000000000
0.100000000000000
-
X
false
-
Rotation:
false
-
Z
false
-
Rotation angle (in degree) around z-axis.
2
-360.000000000000000
360.000000000000000
0.100000000000000
-
Translation (in mm) in direction of the x-axis.
2
-1000.000000000000000
1000.000000000000000
0.100000000000000
-
Scaling:
false
-
Scaling factor for selected fiber bundle along the x-axis.
0.010000000000000
10.000000000000000
0.010000000000000
1.000000000000000
-
Scaling factor for selected fiber bundle along the y-axis.
0.010000000000000
10.000000000000000
0.010000000000000
1.000000000000000
-
Scaling factor for selected fiber bundle along the z-axis.
0.010000000000000
10.000000000000000
0.010000000000000
1.000000000000000
-
false
QCommandLinkButton:disabled {
border: none;
}
Copy Bundles
:/QmitkDiffusionImaging/general_icons/copy2.ico:/QmitkDiffusionImaging/general_icons/copy2.ico
-
false
QCommandLinkButton:disabled {
border: none;
}
Transform Selection
:/QmitkDiffusionImaging/general_icons/refresh.ico:/QmitkDiffusionImaging/general_icons/refresh.ico
-
false
QCommandLinkButton:disabled {
border: none;
}
Join Bundles
:/QmitkDiffusionImaging/general_icons/plus.ico:/QmitkDiffusionImaging/general_icons/plus.ico
-
If checked, the fiducials belonging to the modified bundle are also modified.
Include Fiducials
true
m_CircleButton
m_FlipButton
m_RealTimeFibers
m_AdvancedOptionsBox
m_DistributionBox
m_VarianceBox
m_FiberDensityBox
m_FiberSamplingBox
m_TensionBox
m_ContinuityBox
m_BiasBox
m_GenerateFibersButton
m_ConstantRadiusBox
m_NumBundlesBox
m_MinDensityBox
m_MaxDensityBox
m_MinTwistBox
m_MaxTwistBox
m_CurvyMinBox
m_CurvyMaxBox
m_SizeMinBox
m_SizeMaxBox
m_StepSizeMinBox
m_StepSizeMaxBox
m_VolumeSizeX
m_VolumeSizeY
m_VolumeSizeZ
m_RandomPhantomButton
m_XrotBox
m_YrotBox
m_ZrotBox
m_XtransBox
m_YtransBox
m_ZtransBox
m_XscaleBox
m_YscaleBox
m_ZscaleBox
m_TransformBundlesButton
m_CopyBundlesButton
m_JoinBundlesButton
m_IncludeFiducials
m_SaveParametersButton
m_LoadParametersButton
m_FidAxis1
m_FidPosY
m_FidPosZ
m_FidAxis2
m_FidTwist
m_AlignOnGrid
m_FidPosX
diff --git a/PythonRequirements.txt b/PythonRequirements.txt
index b2d3e9d..a2443a8 100644
--- a/PythonRequirements.txt
+++ b/PythonRequirements.txt
@@ -1,9 +1,9 @@
numpy
SimpleITK
dipy
sklearn
https://github.com/MIC-DKFZ/batchgenerators/archive/v0.19.5.zip
-https://github.com/MIC-DKFZ/TractSeg/archive/v2.1.1.zip
+https://github.com/MIC-DKFZ/TractSeg/archive/master.zip
https://github.com/MIC-DKFZ/HD-BET/archive/master.zip
torch
torchvision
diff --git a/README.md b/README.md
index 5429a84..1e2d060 100644
--- a/README.md
+++ b/README.md
@@ -1,239 +1,239 @@
MITK Diffusion
==============
Copyright © German Cancer Research Center (DKFZ), [Division of Medical Image Computing (MIC)](https://www.dkfz.de/en/mic/index.php). Please make sure that your usage of this code is in compliance with the code [license](https://github.com/MIC-DKFZ/MITK-Diffusion/blob/master/LICENSE.txt).
The MITK Diffusion application [[1,2]](#References) offers a selection of image analysis algorithms for the processing of diffusion-weighted MR images. It encompasses the research of the Division of Medical Image Computing at the German Cancer Research Center (DKFZ).
* [Downloads](#Downloads)
* [Requirements](#Requirements)
* [Features](#Features)
* [Related Links](#Related-Links)
* [Image Gallery](#Image-Gallery)
* [Building MITK Diffusion from source](#Building-MITK-Diffusion-from-source)
* [User Manual](http://docs.mitk.org/nightly/org_mitk_gui_qt_diffusionimaging.html)
* [Report a Bug](https://phabricator.mitk.org/maniphest/task/edit/form/29/)
* [References](#References)
* [Contact](#Contact)
## Downloads
**Nightly Ubuntu and Windows installers: [ftp://ftp.dkfz-heidelberg.de/outgoing/MitkDiffusion](https://bit.ly/2S1QfC8)**
Please also have a look at the [requirements](#Requirements) for running MITK Diffusion with all its features successfully!
The installers come as executable setup wizards that install MITK Diffusion on your system or alternatively as simple .tar.gz or .zip archive where you can execute MITK Diffusion and the command line apps "manually". Should there be no new installer for a while, please [contact](#Contact) us and report the issue.
If you encounter any bugs, please report them in our [bugtracking](https://phabricator.mitk.org/maniphest/task/edit/form/29/) system or use the [MITK-users mailing list](http://mitk.org/wiki/MITK_Mailinglist). We are grateful for any feedback!
### Requirements
* Install Python 3.X: `sudo apt install python3 python3-pip` (Ubuntu) or from https://www.python.org/downloads/windows/ (Windows)
* Download our Python requirements file: [PythonRequirements.txt](https://github.com/MIC-DKFZ/MITK-Diffusion/tree/master/PythonRequirements.txt)
* Install the Python requirements: `pip3 install -r PythonRequirements.txt`
* If your are behind a proxy use `pip3 --proxy install -r PythonRequirements.txt`
**For Windows users**:
MITK Diffusion requires the Microsoft Visual C++ 2017 Redistributable to be installed on the system. The MITK Diffusion installer automatically installs this redistributable for you if not already present on the system, but it needs administrative privileges to do so. So to install the redistributable, **run the MITK Diffusion installer as administrator**.
## Features
**Support for most established image formats**
* Images: DICOM, NIFTI, NRRD (peak and SH images compatible with MRtrix)
* Tractograms: fib/vtk, tck and trk.
**Image preprocessing**
* Registration
* Head-motion correction
* Denoising
* Skull stripping and brain mask segmentation (Linux only)
* Resampling, cropping, flipping and merging
* Header modifications
* Single volume extraction
**Diffusion gradient/b-value processing**
* b-value rounding
* Gradient direction flipping
* Gradient direction subsampling
* Averaging of gradient directions/volumes
* Gradient direction and b-value visualization
**ODF reconstruction and signal modelling**
* Tensor and Q-ball reconstruction
* Other reconstructions via Dipy wrapping (CSD, 3D SHORE, SFM) (Linux only)
* ODF peak calculation
* MRtrix or camino results can be imported
**Quantification of diffusion-weighted/tensor/ODF images**
* Intravoxel Incoherent Motion (IVIM) and diffusion kurtosis analysis
* Calculation of many other derived indices such as ADC, MD, GFA, FA, RA, AD, RD
* Image statistics
**Segmentation**
* Automatic white matter bundle segmentation (TractSeg) [[3]](#References) (Linux only)
* Automatic brain mask segmentation (Linux only)
* Manual image segmentation and operations on segmentations
* SOON: automatic brain tissue segmentation
**Fiber tractography**
* Global tractography [[4]](#References)
* Streamline tractography
* Interactive (similar to [[5]](#References)) or seed image based
* Deterministic or probabilistic
* Peak, ODF, tensor and raw dMRI based. The latter one in conjunction with machine learning based tractography [[6]](#References)
* Various possibilities for anatomical constraints.
* Tractography priors in form of additional peak images, e.g. obtained using TractSeg
**Fiber processing**
* Tract dissection (parcellation or ROI based)
* Tract filtering by
* length
* curvature
* direction
* weight
* density
* Tract resampling and compression
* Tract transformation
* Mirroring
* Rotating and translating
* Registration (apply transform of previously performed image registration)
* Tract coloring
* Curvature
* Length
* Weight
* Scalar map (e.g. FA)
* Other operations
* Join
* Subtract
* Copy
* Fiber clustering [[7]](#References)
* Fiber fitting and weighting similar to SIFT2 and LiFE [[8,9]](#References)
* Principal direction extraction (fibers --> peaks)
* Tract derived images:
* Tract density images
* Tract endpoint images
* Tract envelopes
**Fiberfox dMRI simulations** [[10]](#References)
* Multi-compartment signal modeling
* Simulation of the k-space acquisition including
* Compartment specific relaxation effects
* Artifacts such as noise, spikes, ghosts, aliasing, distortions, signal drift, head motion, eddy currents and Gibbs ringing
* Definition of important acquisition parameters such as bvalues and gradient directions, TE, TR, dwell time, partial Fourier, ...
* Manual definition of fiber configurations, e.g. for evaluation purposes
* Automatic generation of random fiber configurations
**Other features**
* Brain network statistics and visualization (connectomics)
* Interactive Python console (Linux only)
* Integrated screenshot maker
* Command line tools for most functionalities
## Related Links
* Great python package for logging your (MITK) command line experiments:
* https://github.com/MIC-DKFZ/cmdint
* `pip3 install cmdint`
* TractSeg reference data of 72 semiautomatically defined bundles in 105 HCP subjects: https://zenodo.org/record/1285152
* TractSeg python package: https://github.com/MIC-DKFZ/TractSeg
* Simulated dMRI images and ground truth of random fiber phantoms in various configurations: https://doi.org/10.5281/zenodo.2533250
* ISMRM 2015 Tractography Challenge Data: https://doi.org/10.5281/zenodo.572345 & https://doi.org/10.5281/zenodo.1007149
## Image Gallery
 Screenshot of the MITK Diffusion Welcome Screen
 Scalar map visualization
 Tensor Visualization
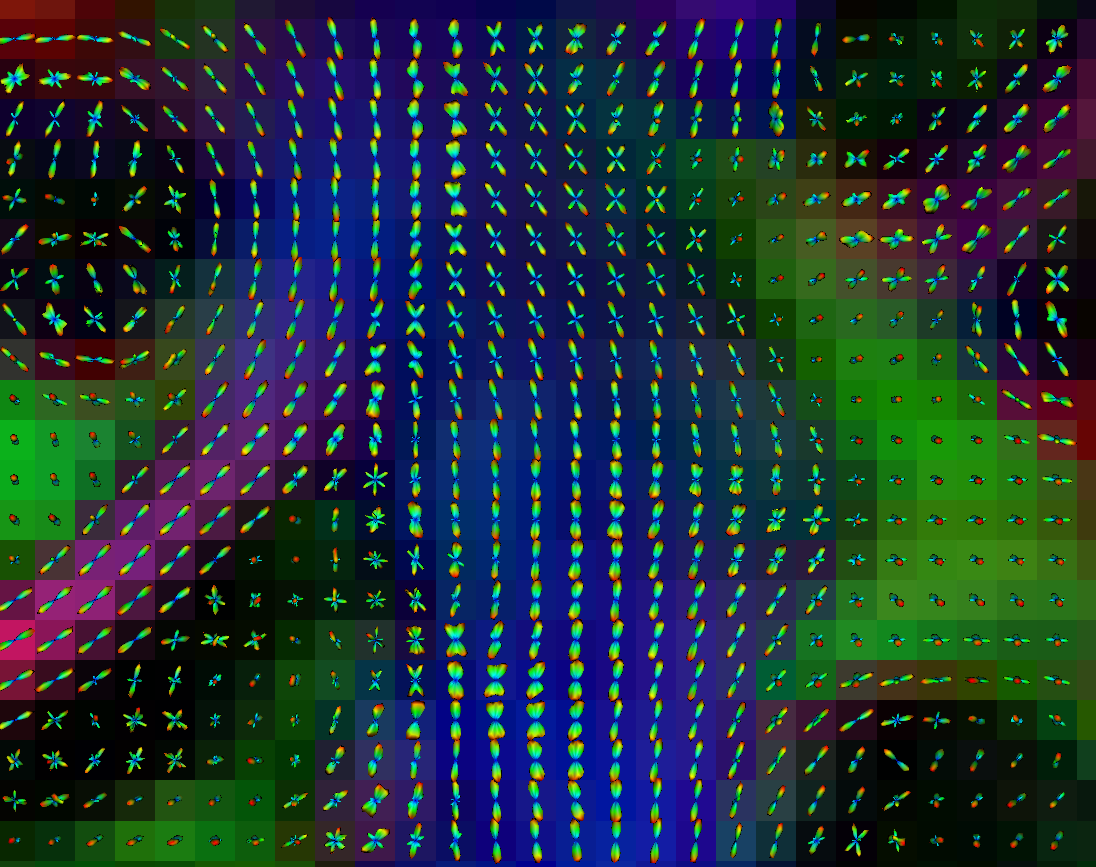 ODF visualization
 Peak visualization (uniform white coloring)
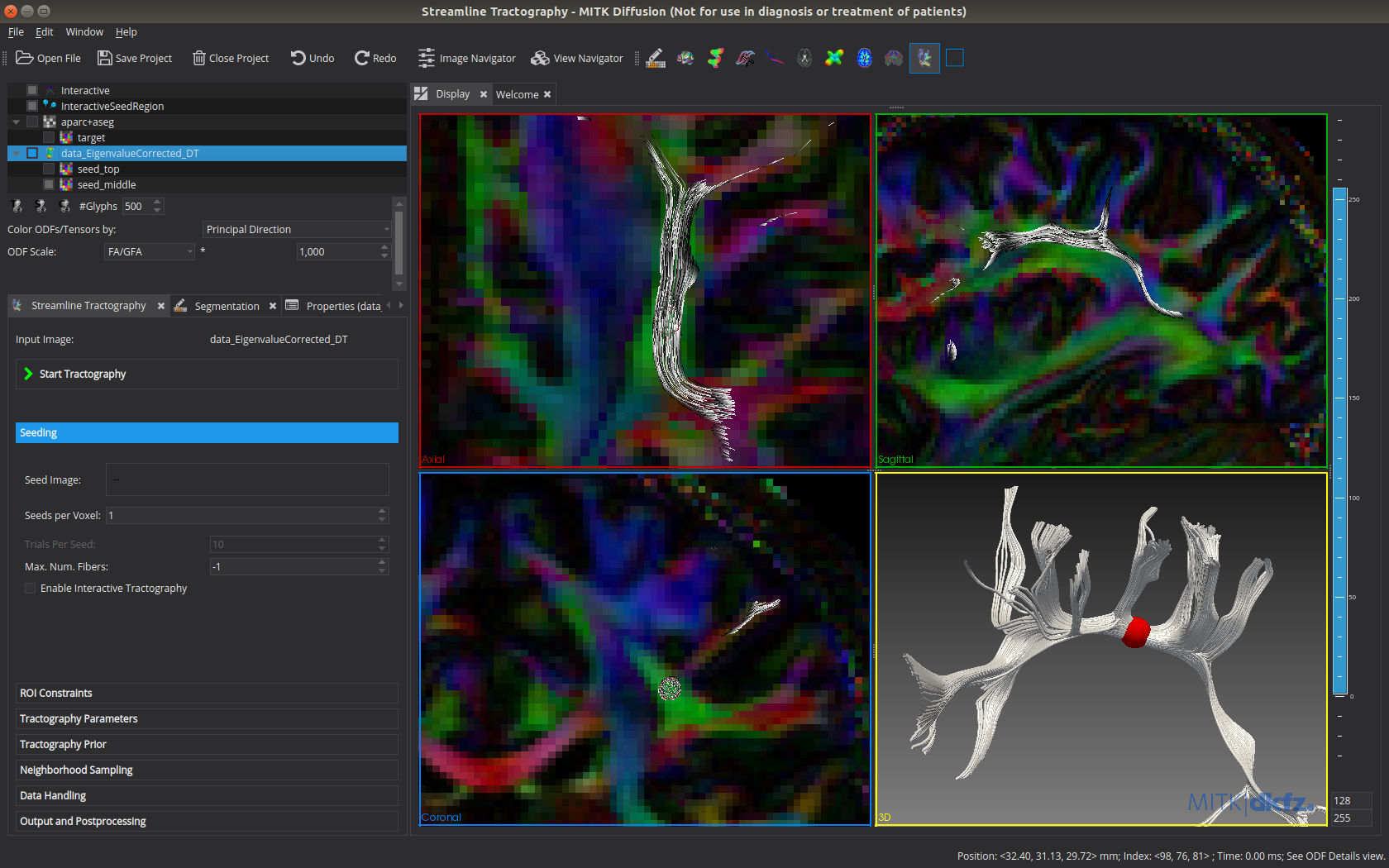 Interactive tractography in MITK Diffusion. The tractogram updates automatically on parameter change and movement of the spherical seed region.
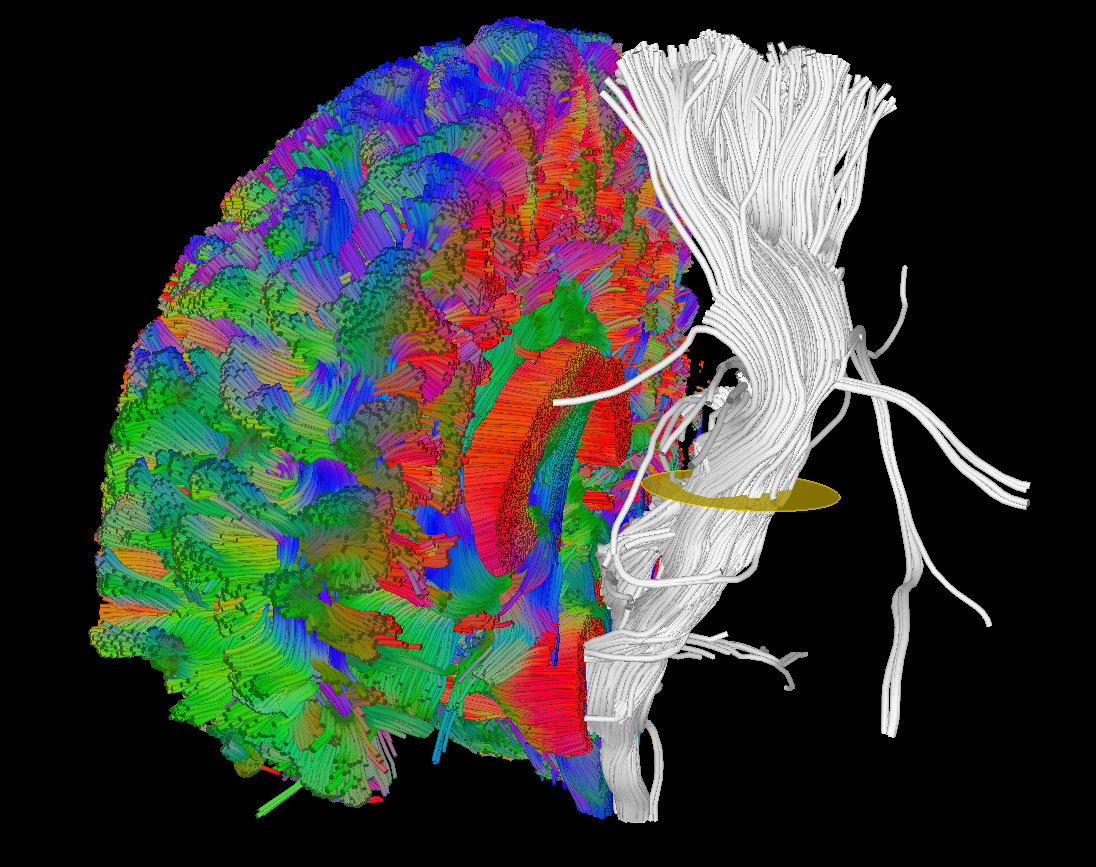 Tract dissection using manually drawn ROIs.
 Automatic streamline weighting (similar to SIFT2 or LiFE)
 Illustration of the dMRI phantom simulation process using Fiberfox.
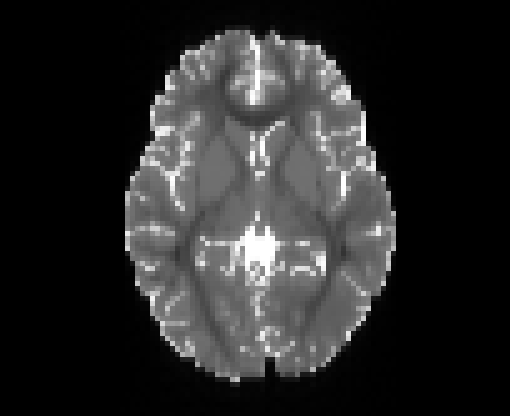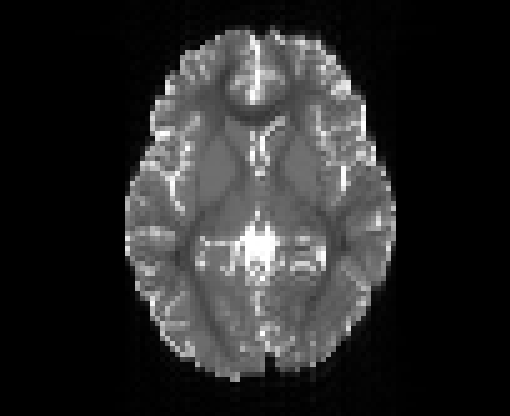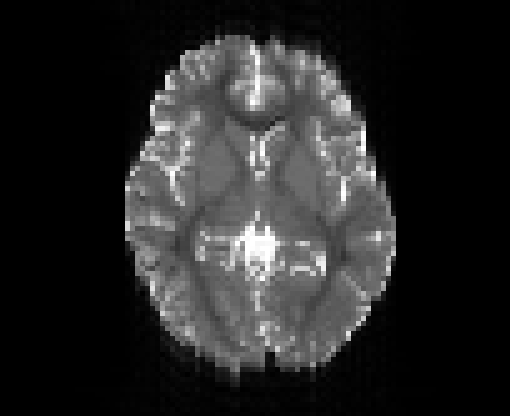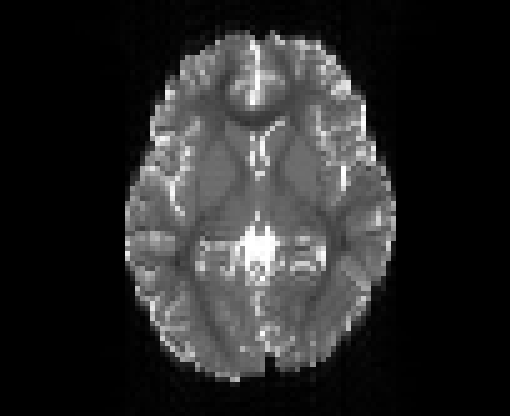
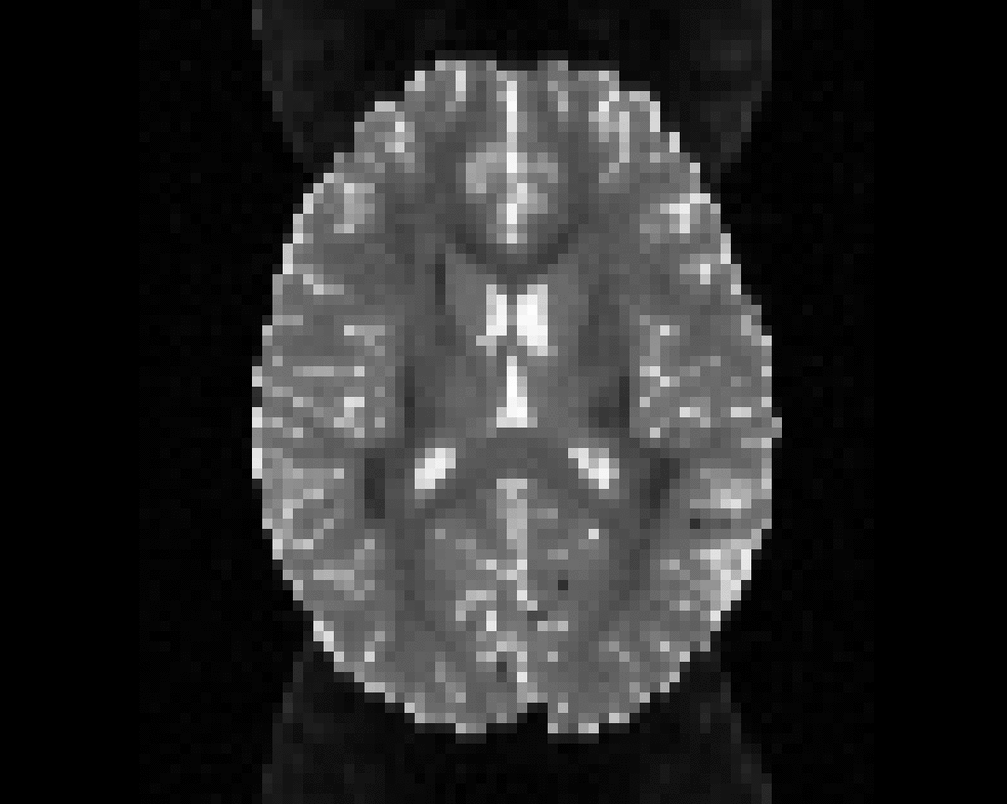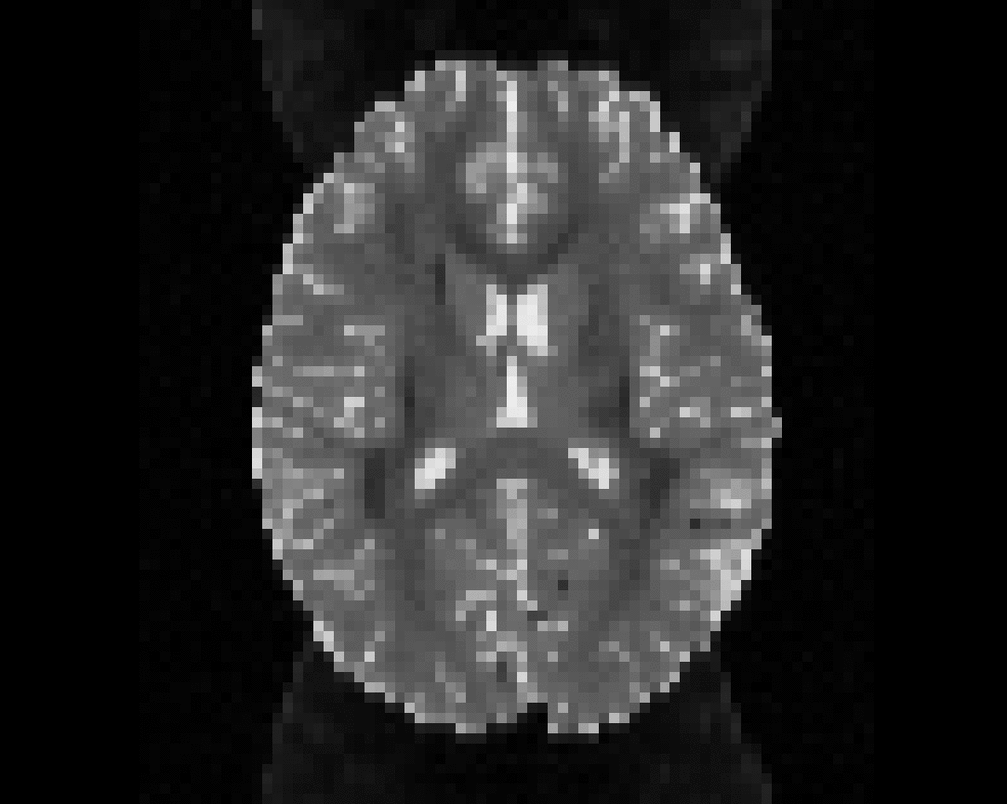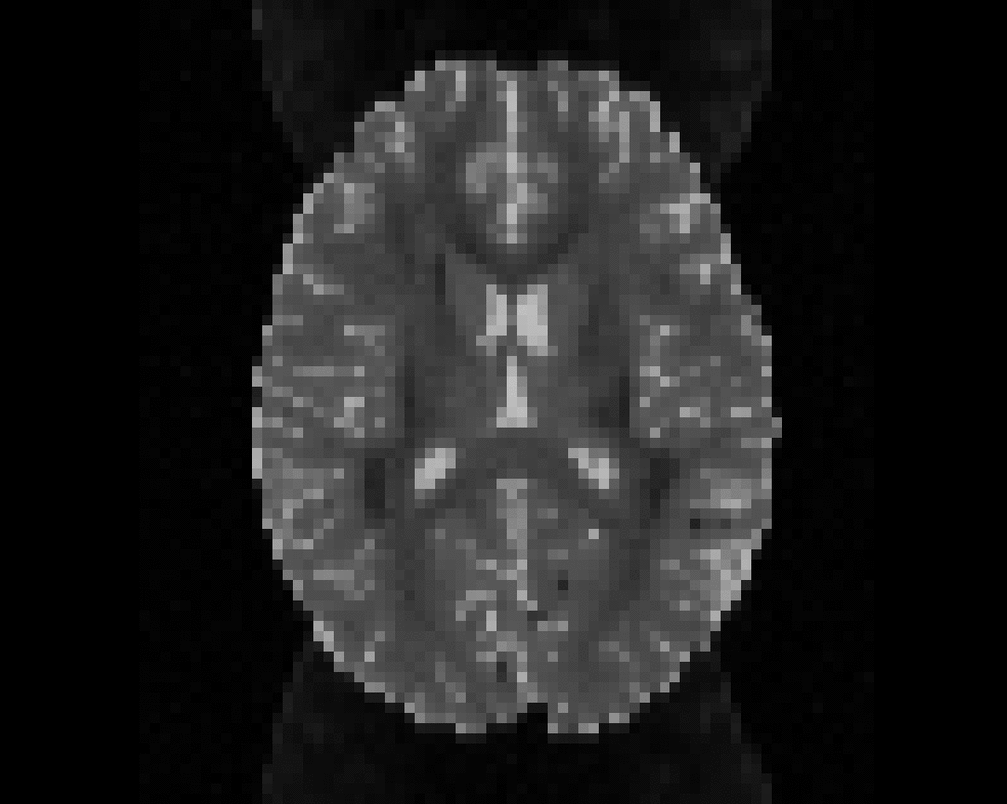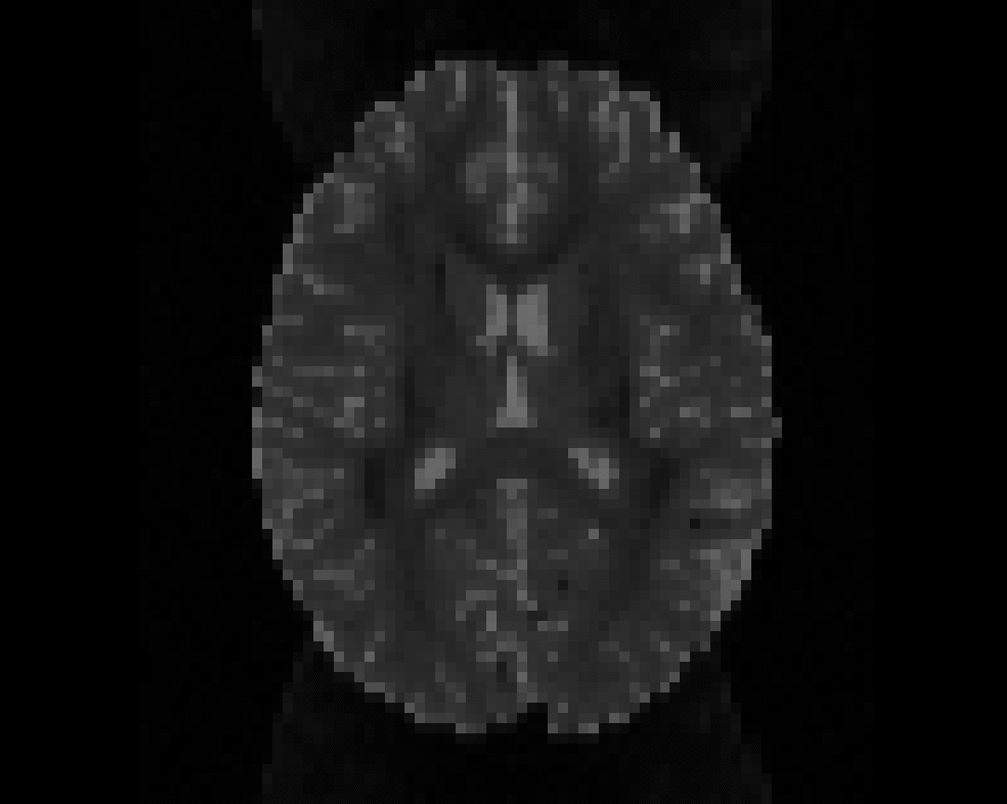
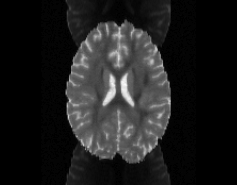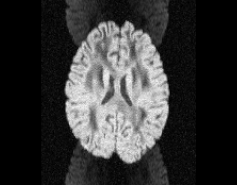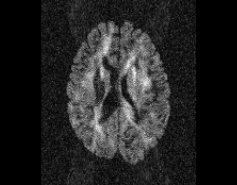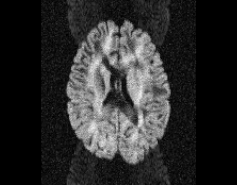
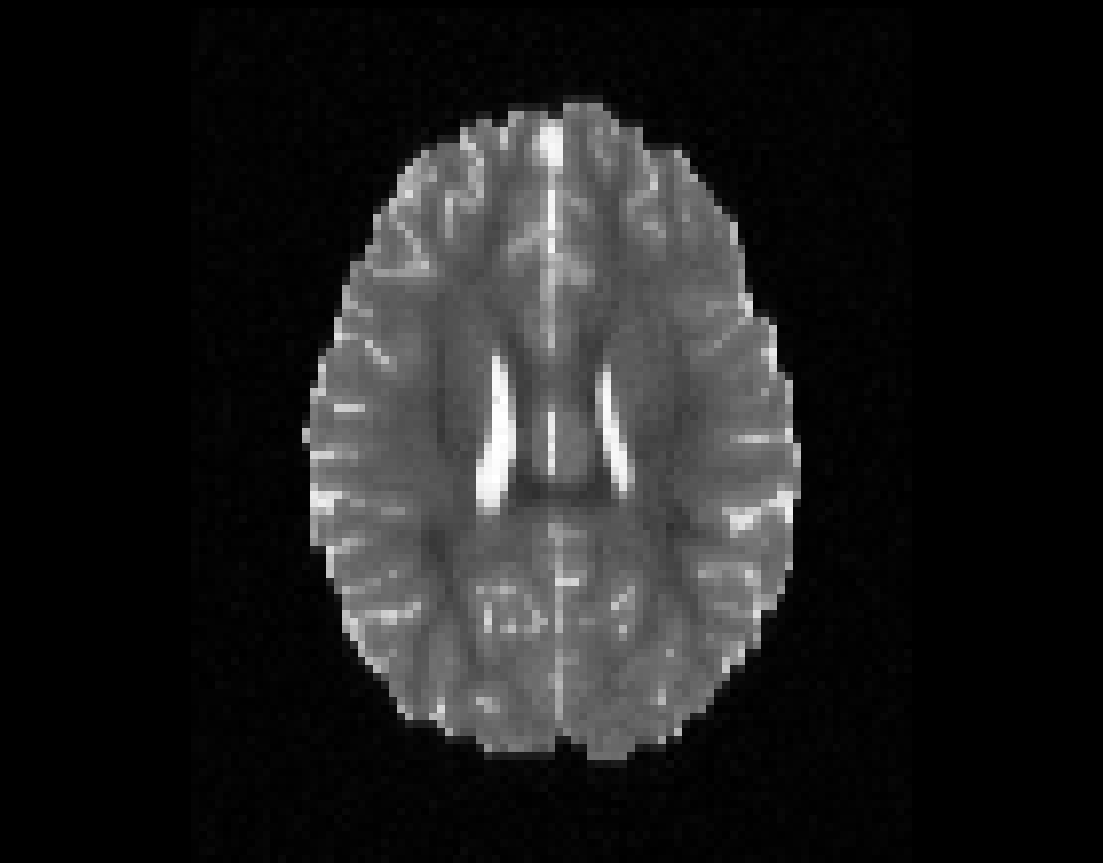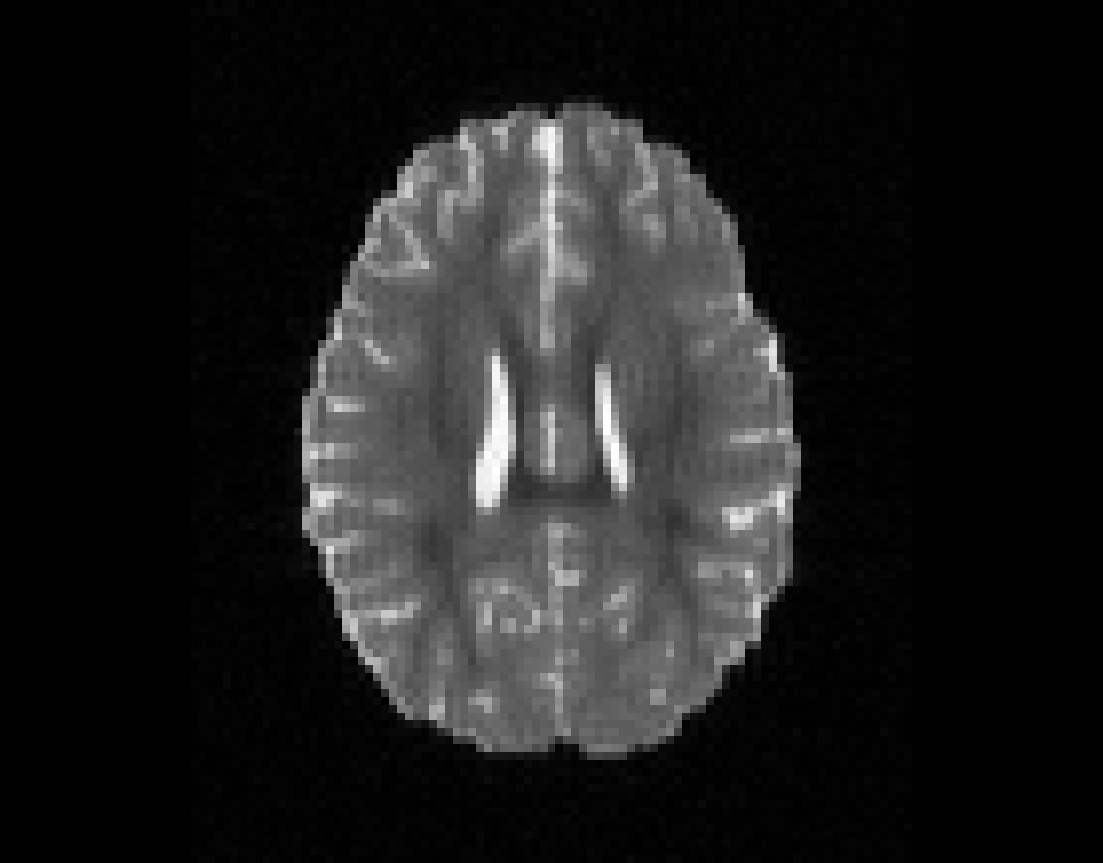
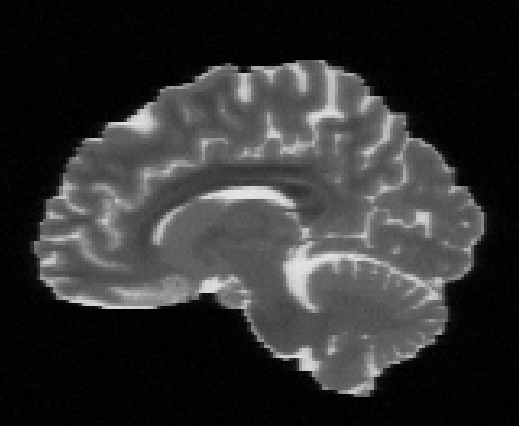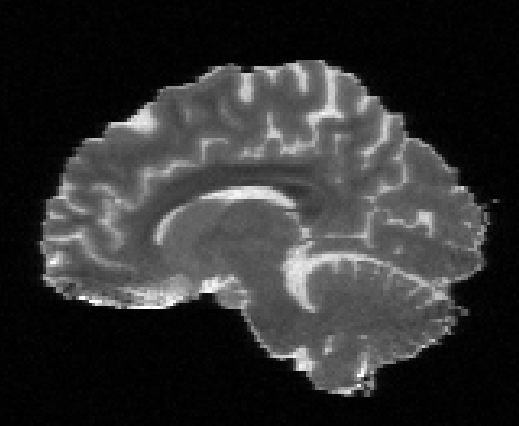 \
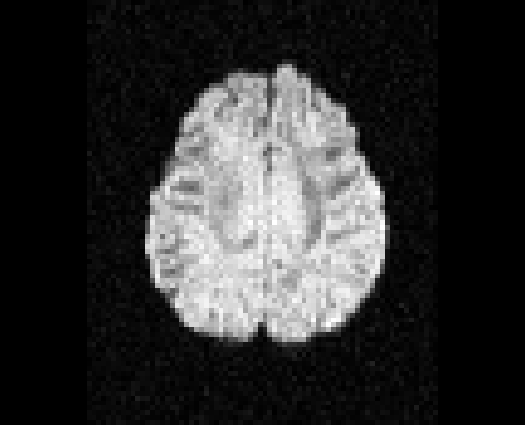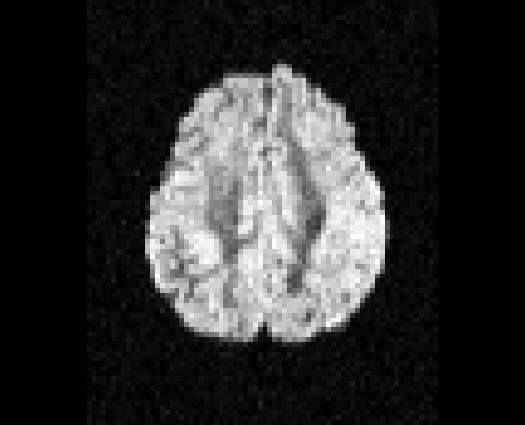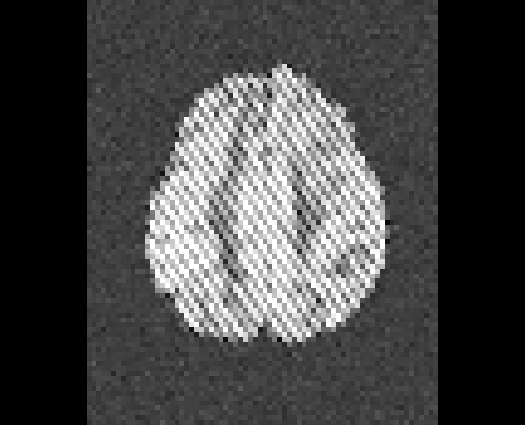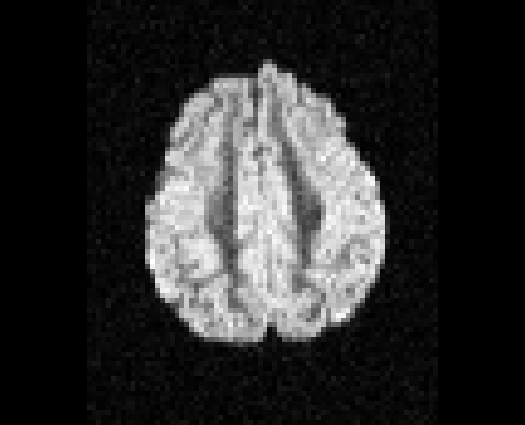
Illustration of simulated dMRI images with various artifacts (a bit excessive for illustration purposes): eddy current distortions (1), motion and spike (2), intensity drift (3), motion, eddy and noise (4), ringing (5), B0 inhomogeneity distortions (6), from left to right.
\
Illustration of simulated dMRI images with various artifacts (a bit excessive for illustration purposes): eddy current distortions (1), motion and spike (2), intensity drift (3), motion, eddy and noise (4), ringing (5), B0 inhomogeneity distortions (6), from left to right.

Automatically generated random fiber configuration for Fiberfox simulations.
## Building MITK Diffusion from source
-* Install [Qt](https://www.qt.io/ Qt) on your system (>= 5.11.1).
+* Install [Qt](https://www.qt.io/) on your system (>= 5.12.6).
* Clone MITK from [github](https://github.com/MIC-DKFZ/MITK-Diffusion.git) using [Git version control](https://git-scm.com/).
* Clone MITK Diffusion from [github](https://github.com/MITK/MITK.git).
* Configure the MITK Superbuild using [CMake](https://cmake.org/) (>= 3.14.5).
* Choose the MITK source code directory and an empty binary directory.
* Click "Configure".
* Set the option MITK_EXTENSION_DIRS to "/path/to/my/mitk-diffusion-repository".
* Click "Configure".
* Set the option MITK_BUILD_CONFIGURATION to "DiffusionRelease".
* Click "Generate".
* macOS specifics:
* Use python 3.**6**, since python 3.**7** leads to build errors on macOS.
* The cmake variables for python 3 might need to be set manually. It is probably enough to specify PYTHON_EXECUTABLE.
* Openmp needs to be installed manually since it is not included in apple clang anymore: "brew install libomp" should do the trick. It might be necessary to set the corresponding make variables later in the MITK build manually:
* OpenMP_CXX_FLAGS: -Xpreprocessor -fopenmp -I"/path/to/python3/includes/"
* OpenMP_C_FLAGS: -Xpreprocessor -fopenmp -I"/path/to/python3/includes/"
* OpenMPCXX_LIB_NAMES: libomp
* OpenMPC_LIB_NAMES: libomp
* OpenMP_libomp_LIBRARY: /path/to/libomp.dylib
* Build the project
* Linux/maxOS: Open a console window, navigate to the build folder and type "make -j8" (optionally supply the number threads to be used for a parallel build with -j).
* Windows (requires visual studio): Open the MITK Superbuild solution file and build all projects.
* The build may take some time and should yield the binaries in "your_build_folder/MITK-build/bin"
More detailed build instructions can be found in the [documentation](http://docs.mitk.org/nightly/BuildInstructionsPage.html).
Continuous integration: http://cdash.mitk.org/index.php?project=MITK&display=project
## References
All publications of the Division of Medical Image Computing can be found [https://www.dkfz.de/en/mic/publications/ here].
[1] Fritzsche, Klaus H., Peter F. Neher, Ignaz Reicht, Thomas van Bruggen, Caspar Goch, Marco Reisert, Marco Nolden, et al. “MITK Diffusion Imaging.” Methods of Information in Medicine 51, no. 5 (2012): 441.
[2] Fritzsche, K., and H.-P. Meinzer. “MITK-DI A New Diffusion Imaging Component for MITK.” In Bildverarbeitung Für Die Medizin, n.d.
[3] Wasserthal, Jakob, Peter Neher, and Klaus H. Maier-Hein. “TractSeg - Fast and Accurate White Matter Tract Segmentation.” NeuroImage 183 (August 4, 2018): 239–53.
[4] Neher, P. F., B. Stieltjes, M. Reisert, I. Reicht, H.P. Meinzer, and K. Maier-Hein. “MITK Global Tractography.” In SPIE Medical Imaging: Image Processing, 2012.
[5] Chamberland, M., K. Whittingstall, D. Fortin, D. Mathieu, und M. Descoteaux. „Real-time multi-peak tractography for instantaneous connectivity display“. Front Neuroinform 8 (2014): 59. doi:10.3389/fninf.2014.00059.
[6] Neher, Peter F., Marc-Alexandre Côté, Jean-Christophe Houde, Maxime Descoteaux, and Klaus H. Maier-Hein. “Fiber Tractography Using Machine Learning.” NeuroImage. Accessed July 17, 2017. doi:10.1016/j.neuroimage.2017.07.028.
[7] Garyfallidis, Eleftherios, Matthew Brett, Marta Morgado Correia, Guy B. Williams, and Ian Nimmo-Smith. “QuickBundles, a Method for Tractography Simplification.” Frontiers in Neuroscience 6 (2012).
[8] Smith, Robert E., Jacques-Donald Tournier, Fernando Calamante, and Alan Connelly. “SIFT2: Enabling Dense Quantitative Assessment of Brain White Matter Connectivity Using Streamlines Tractography.” NeuroImage 119, no. Supplement C (October 1, 2015): 338–51.
[9] Pestilli, Franco, Jason D. Yeatman, Ariel Rokem, Kendrick N. Kay, and Brian A. Wandell. “Evaluation and Statistical Inference for Human Connectomes.” Nature Methods 11, no. 10 (October 2014): 1058–63.
[10] Neher, Peter F., Frederik B. Laun, Bram Stieltjes, and Klaus H. Maier-Hein. “Fiberfox: Facilitating the Creation of Realistic White Matter Software Phantoms.” Magnetic Resonance in Medicine 72, no. 5 (November 2014): 1460–70. doi:10.1002/mrm.25045.
## Contact
If you have questions about the application or if you would like to give us feedback, don't hesitate to contact us using [our mailing list](http://mitk.org/wiki/MITK_Mailinglist) or, for questions that are of no interest for the community, [directly](https://www.dkfz.de/en/mic/team/people/Peter_Neher.html).





 \
Illustration of simulated dMRI images with various artifacts (a bit excessive for illustration purposes): eddy current distortions (1), motion and spike (2), intensity drift (3), motion, eddy and noise (4), ringing (5), B0 inhomogeneity distortions (6), from left to right.
\
Illustration of simulated dMRI images with various artifacts (a bit excessive for illustration purposes): eddy current distortions (1), motion and spike (2), intensity drift (3), motion, eddy and noise (4), ringing (5), B0 inhomogeneity distortions (6), from left to right.